Overview
DisCVR: Rapid viral diagnosis from HTS data
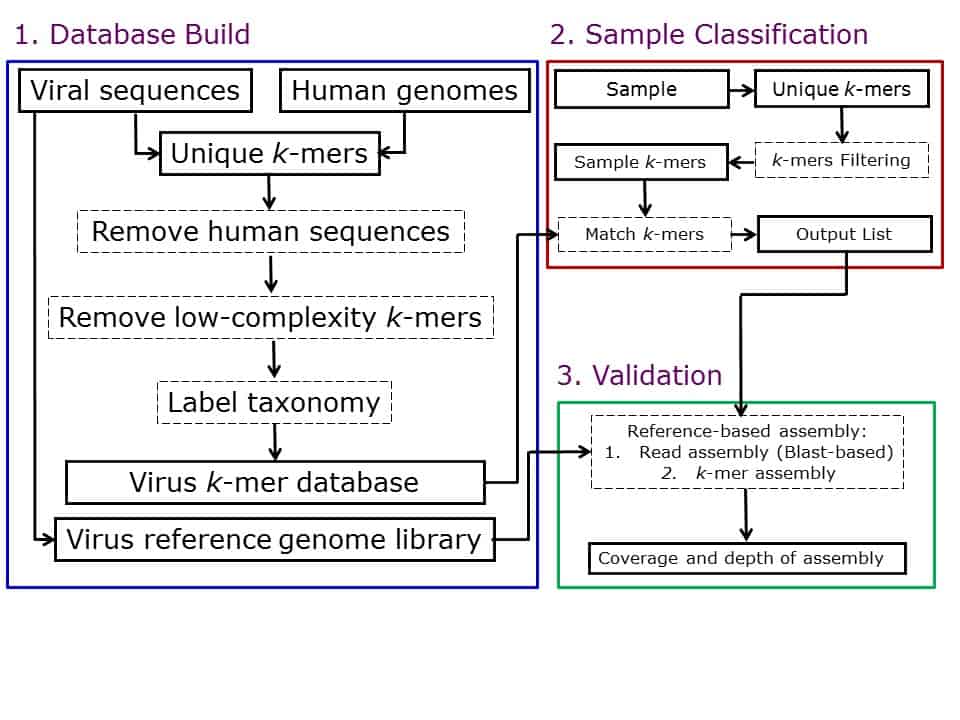
DisCVR is a computer program which allows diagnosticians to detect known human viruses in clinical samples from High-Throughput Sequencing (HTS) data. It works by creating a database of short nucleotide sequences, called k-mers, which are extracted from viral genomes. k-mers of 22 bases in length are generated by sliding a window along a sequence, 1 nucleotide base at a time. Only unique k-mers from a set of viruses are included in the database and assigned taxonomic labels. To investigate a patient sample sequenced using HTS, the database is queried to find exact matches with k-mers from each read in the sample. A list of all viruses found in the sample is shown and reference-based assembly can be used to show depth and coverage of data in relation to a reference genome, in order to assess significance of matches.
DisCVR is a fast and accurate tool designed to analyse HTS data and validate the results interactively on computers with limited resources. At present, DisCVR is a human viral diagnostic tool, but it could be extended to include non-viral human pathogens as well as pathogens of other hosts.
Test samples
- Three respiratory samples are provided to test run DisCVR. They can be dowloaded here (166Mb).
References
- Kanalyze: A Fast Versatile Pipelined K-mer Toolkit, Bioinformatics 30(14) (2014) 2070-2072
- TANOTI: A rapid BLAST-guided read mapper for small, divergent genomes.(manuscript communicated)
- JFreechart: Free Java chart library
Contact
Please send your comments, suggestions or bug reports to Joseph Hughes.
