Intro
-
Tanoti: (तनोति) (Sanskrit): Weave, Manifest, Accomplish, Augment, Render
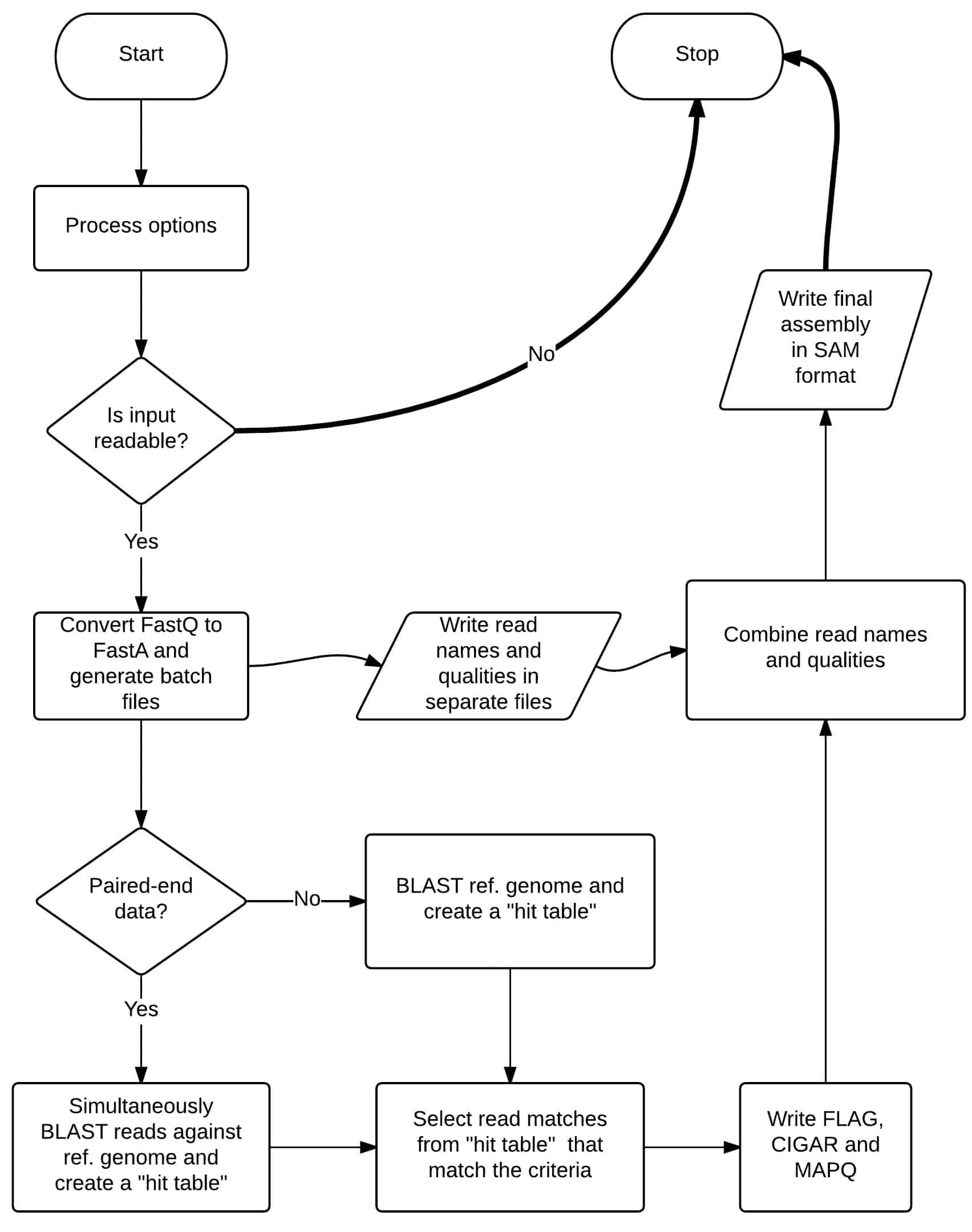
Tanoti is a BLAST guided reference based short read aligner. It is developed for maximising alignment in highly variable next generation sequence data sets (Illumina).
Dynamic memory management and batch processing make Tanoti consume very little memory and processing power. Tanoti’s read alignment performance is superior to BWA and Bowtie in our comparisons on small viral genomes, giving greater depth with highly variable reads without losing accuracy of alignment.
Download
- Linux (Intel) Click here
- Mac OSX Click here
- (Source code available on GitHub)
Reference
TANOTI: a rapid BLAST-guided read mapper for small, divergent genomes.(manuscript communicated)
Usage
Paired end reads:
tanoti –r reference.fa –i input1.fq input2.fq -o output.sam –p 1
Single end reads:
tanoti –r reference.fa –i input1.fq -o output.sam
Where:
-r : reference genome(s) in FASTA format
-i : input files in FASTQ format
-o : output file name (default: FinalAssembly.sam)
-p : paired-end reads 1/0 (default 0)
Optional parameters:
-m : minimum match percentage of a read (default 50%)
-u : include unmapped reads in the output 1/0 (default 0)
-t : Keep temporary files
-P : number of parallel BLAST searches (default 2/single end and 4/paired end. Don’t change this value if you are unsure of it
-h : Help
Reference
TANOTI: a rapid BLAST-guided read mapper for small, divergent genomes.(manuscript communicated)
Contact
Please send your comments, suggestions or bug reports to Dr. Sreenu Vattipally
