How to Import data for libraries with index tags into BaseSpace
- Post by: Gavin Wilkie
- April 4, 2016
- No Comment
In this blog we describe how to import lists of sample data with defined index tags into BaseSpace, and provide templates for TruSeqLT and TruSeqHT libraries. We have found this saves a lot of time and eliminates errors associated with manual entry.
The Illumina NextSeq500 sequencer requires all users to complete sample data entry on BaseSpace (Illumina’s cloud-based resource) including sample names, species, project names, index tags and sample pools. Whilst there are many advantages to having this data in the cloud, the BaseSpace interface is not always the most convenient or user-friendly system for data entry and management.
Our experience has been that for large projects with many samples, it is impractical to use the manual method of entering sample names in the ‘Biological Samples’ tab, then individually assigning an index tag in the ‘Libraries’ tab by dragging each sample onto an image of a 96-well plate of barcodes. To make matters worse, BaseSpace always mixes up the order of the samples (even if they are named 1-96), so it becomes all too easy to make an error when faced with a long list of sample names in a random order that each require a tag to be assigned.
It is quite easy to import a csv file created in Excel (or similar) with the sample names, species, project and nucleic acid into the ‘Biological Samples’ tab, and thus avoid a large part of the manual data entry. However this still requires the user to individually assign an index tag to each sample using the cumbersome and error-prone interface pictured below, dragging each sample on the list to the correct well on the index plate.
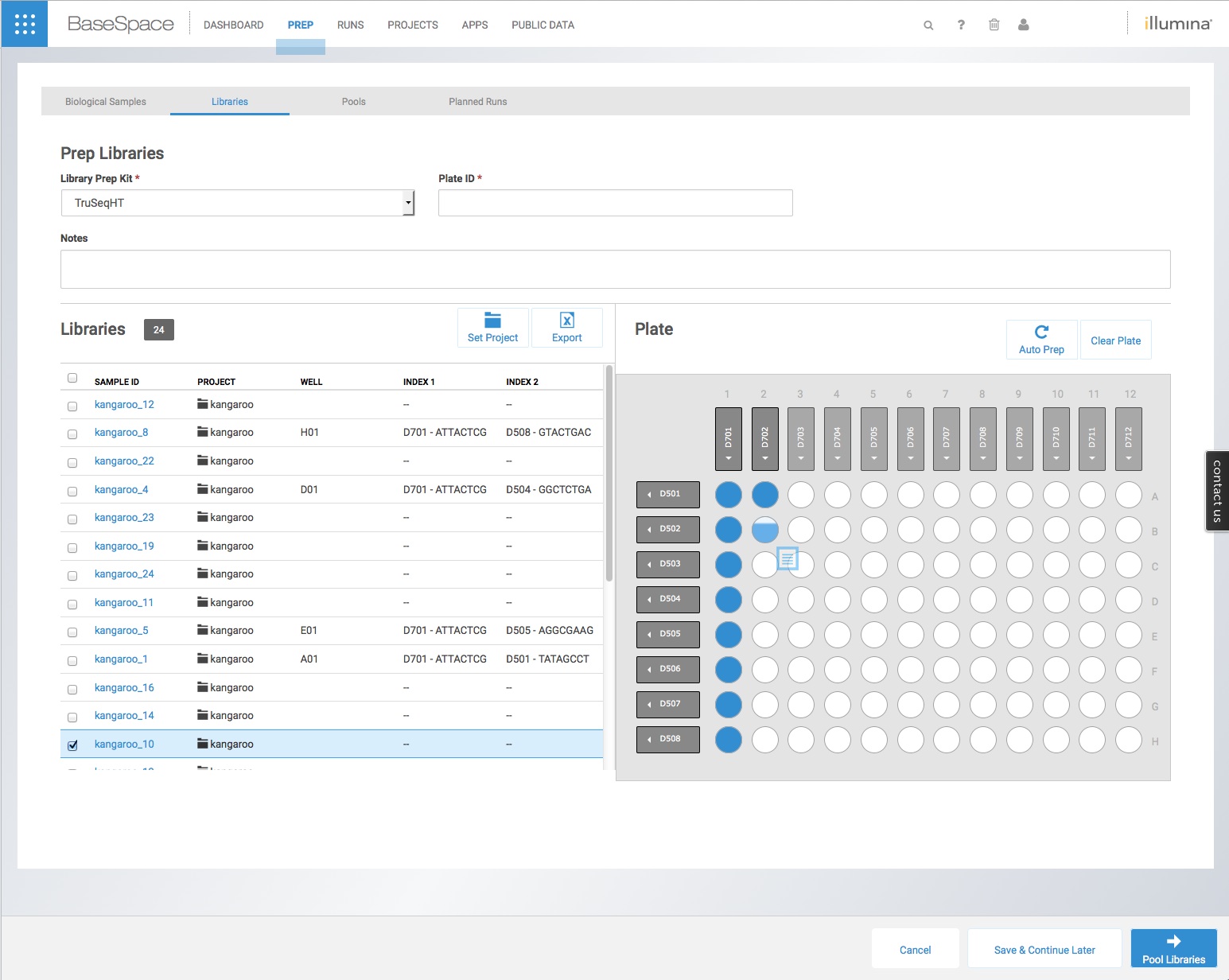
It is possible to avoid this by importing a csv file with the sample names, species, project, nucleic acid, index name and also the index tags into the ‘Libraries’ tab on BaseSpace. However, there is very little guidance on how to do this – and Illumina only provide an example template for libraries made using Nextera XT with none of the sequence tags themselves.
We are mainly using TruSeq indexes, so we have generated our own import templates with all 24 TruSeqLT tags, and all 96 dual-indexed TruSeqHT tags. This took quite a bit of trial and error, plus fetching the sequences of all 216 index tags. We have therefore made our own templates for importing TruSeqLT and TruSeqHT libraries available here for others to use.
Simply open the csv file in Excel (or similar) and insert the names of your own samples in the first two columns. Copy and past the index tags you have used to the correct sample lines (Each sample requires the Well, Index1Name, Index1Sequence,Index2Name and Index2Sequence). Change the name of the ContainerID from ‘Platename’ to your own name and delete any lines you don’t need (e.g. if you have less than 24 or 96 samples). Here we are using the template to import 24 samples called apples 1-24 with TruSeqHT dual tags.If using 96 samples, use this.
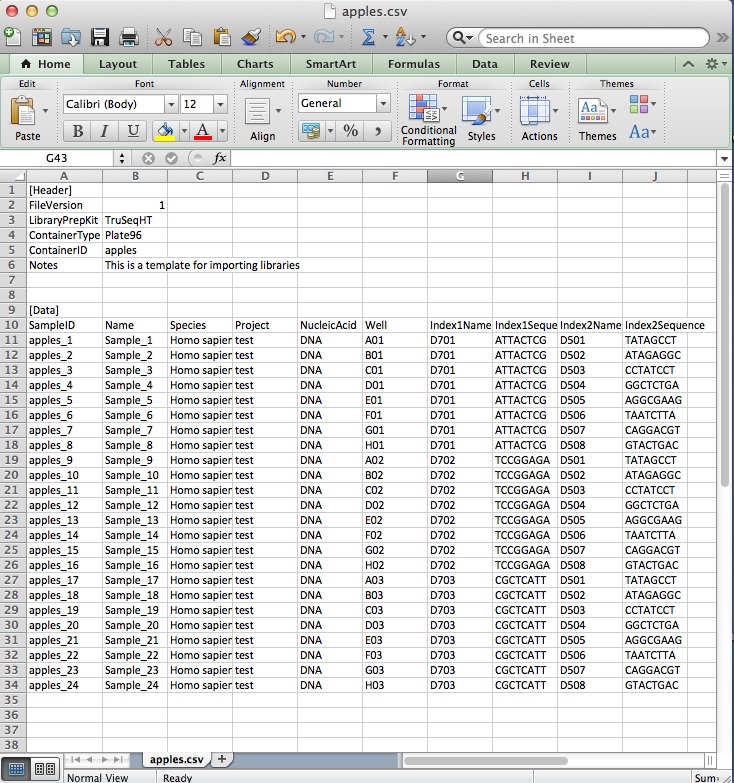
Save the csv file, navigate to the ‘Libraries’ tab in your BaseSpace account and then click the ‘Import’ button on the top-right corner. Choose your csv file, and after a minute you should see your libraries successfully imported with the correct index tags as below, ready to pool for a sequence run.
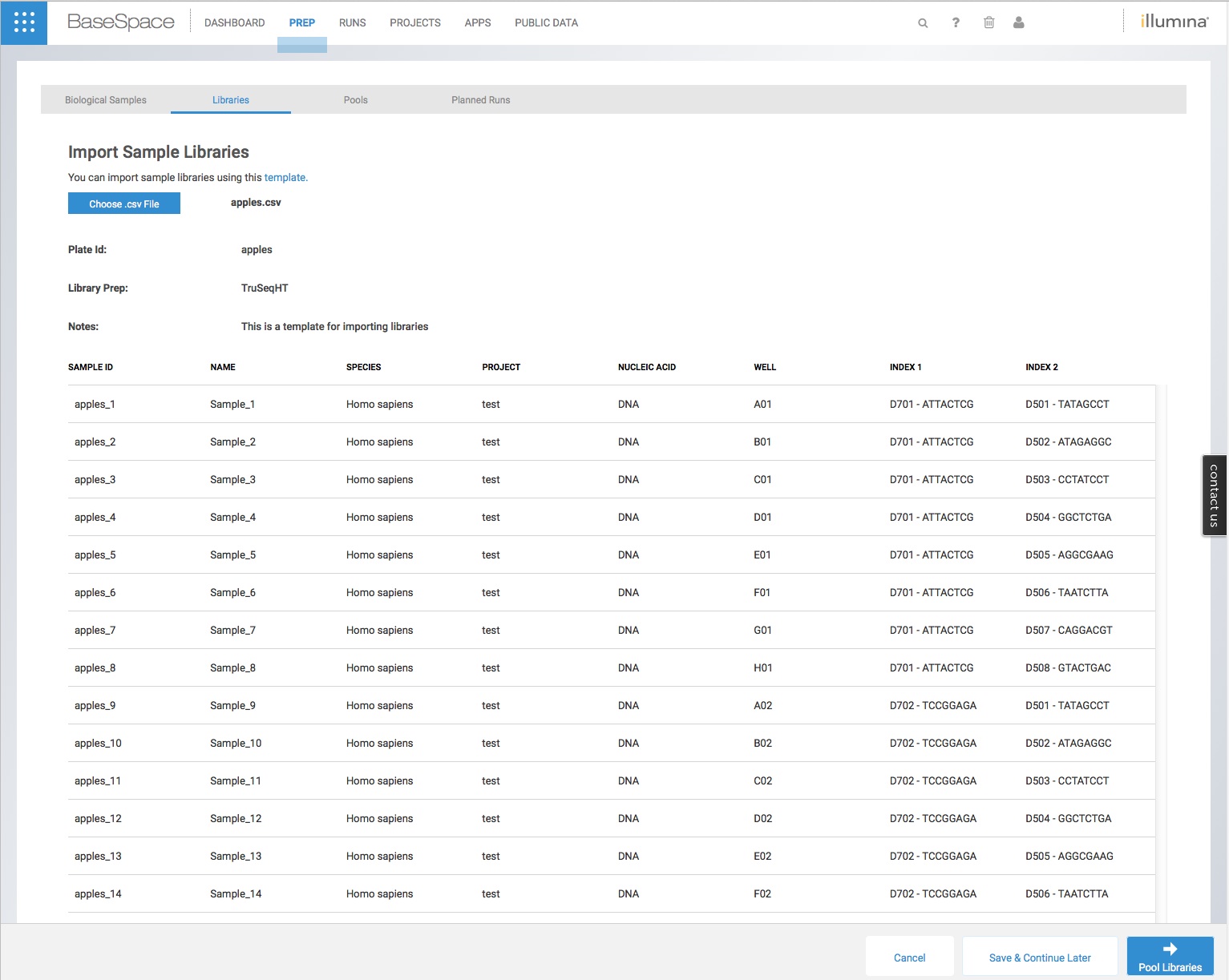
Now, if Illumina would just allow us to import pools of samples we could also avoid having to individually drag each sample into a small dot in the ‘Pools’ tab. This is rather tiresome when there are large numbers of samples in a pool!
Categories: Deep Sequencing, illumina
